Immunohistochemical detection of putative pathogenetic factors in adhesions of infants: a pilot study

All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article or claim that may be made by its manufacturer is not guaranteed or endorsed by the publisher.
Authors
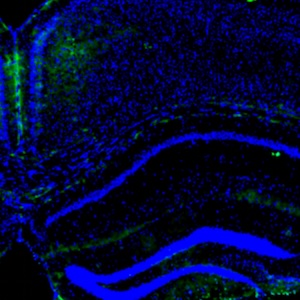
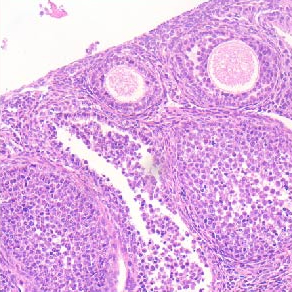
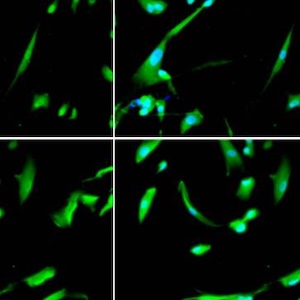
Newborns’ intestinal adhesions have been reported in 4.7% infants who underwent a laparotomy, but adhesions can also appear idiopathically. Etiology and pathogenesis of adhesions is still to be determined, but evidence shows relation to inflammation, formation of fibrin bands, hypoxia and tissue remodelation. Multiple candidate genes have been associated with adhesion development. The aim of this study was to evaluate the appearance of Sonic Hedgehog (SHH), Indian Hedgehog (IHH), Forkhead-box F1 (FOXF1), caudal type homeobox 1 (CDX1), HCLS1-associated protein X-1 (HAX-1), GATA Binding Protein 4 (GATA4) and Granzyme-B (GZMB) proteins in infant adhesions and to describe possible interfactorial correlations. Adhesion affected tissue samples were collected from 14 patients under one year of age that underwent abdominal surgery to treat partial or complete intestinal obstruction. The control group consisted of 6 individuals that had surgical repairment of inguinal hernia. Routine staining and immunohistochemistry were performed. Immunopositive fibroblasts, macrophages, endotheliocytes, smooth muscle myocytes of blood vessel wall and mesotheliocytes were investigated. The relative distribution of all factors was evaluated by the semiquantitative counting method. Statistical analysis was done using non-parametric tests and correlations were calculated based on Spearman’s correlation analysis. A statistically significant decrease was observed for SHH, IHH, FOXF1, GATA4 and partially for GZMB in the adhesion group. There were also decreased HAX-1 and CDX1 immunopositive structures in the adhesion group, however, without any statistical significance. SHH, IHH, FOXF1, GATA4 and GZMB might have a role in adhesion development among infant patients which could suggest a dysregulation of cellular events. Abundance of correlations between the gene protein appearances in different structures indicate the affected blood vessels, fibroblasts and macrophages, however, mesothelium seems not to be the key driver in the morphopathogenesis of adhesion development.
Ethics Approval
this research study was approved by Rīga Stradiņš University Ethical committee on 10.05.2007.CRediT authorship contribution
Arvis Pauliņš, formal analysis, software, visualization, writing - original draft, review and editing. Anna Junga, data curation, investigation, methodology, resources; Māra Pilmane, conceptualization, data curation, investigation, methodology, resources, funding acquisition, project administration, supervision, validation, writing - review and editing.
Data Availability Statement
The data used and analyzed in this research are presented in the results section of this study.
How to Cite

This work is licensed under a Creative Commons Attribution-NonCommercial 4.0 International License.